A List Of Heredity Related Molecules:
It is clear that this is a quite complex issue. In this list there are some of these molecules and there may be more to be added.
As we know some of these are quite well known as DNA, RNA and so on. On the other hand this microenvironment is quite detailed.
DNA and RNA is some of the most very well knowns of these molecules.
There may be a short list of these mollecules as:
“While DNA is the primary genetic material in most organisms, RNA also plays critical roles in gene expression and inheritance. There are different types of RNA molecules involved in various aspects of genetic regulation and information transfer. Here are some key RNA molecules related to heredity:
- Messenger RNA (mRNA): mRNA carries genetic information from DNA to the ribosomes, where it serves as a template for protein synthesis during the process of translation.
- Transfer RNA (tRNA): tRNA molecules bring specific amino acids to the ribosomes during translation, based on the instructions provided by the mRNA sequence.
- Ribosomal RNA (rRNA): rRNA is a crucial component of ribosomes, the cellular structures where proteins are synthesized. It helps in the assembly of the ribosome and facilitates the catalytic functions of protein synthesis.”
So, briefly:
DNA
RNA
Epigenetic Factors
Chromosomes
Transposones
Enzymes
Viral Vectors
Growth Factors
Trascription Factors
Proteins and
Small Molecules

Also a longer and more detailed list may seen and added/list as:
- DNA (Deoxyribonucleic acid): The molecule that carries the genetic instructions for the development, functioning, and reproduction of all living organisms.
- RNA (Ribonucleic acid): A molecule involved in various cellular processes, including gene expression, protein synthesis, and regulation of genetic information.
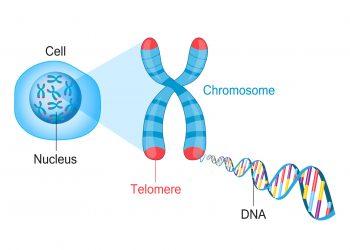
- Chromosomes: Structures within cells that contain DNA and associated proteins, carrying genes and maintaining the stability of genetic material.
- Histones: Proteins that help package DNA into compact structures called nucleosomes and play a role in regulating gene expression.
- Nucleotides: The building blocks of DNA and RNA, consisting of a sugar molecule, a phosphate group, and a nitrogenous base.
- Non-coding DNA: DNA sequences that do not encode proteins but play roles in gene regulation, chromosome structure, and genome stability.
- Transcription factors: Proteins that bind to DNA and regulate the transcription of genes by either activating or repressing their expression.
- RNA polymerase: An enzyme that catalyzes the synthesis of RNA from a DNA template during transcription.
- Proteins: Macromolecules composed of amino acids that perform various functions in cells, including structural support, enzymatic activity, and cellular signaling.
- Enzymes: Proteins that catalyze biochemical reactions in cells, including those involved in DNA replication, repair, and modification.
- Viral vectors: Modified viruses used as vehicles to deliver genetic material into cells for various research or therapeutic purposes.
- Growth factors: Signaling molecules that regulate cell growth, proliferation, and differentiation during development and tissue repair.
- Small molecules: Organic compounds with low molecular weight that can have regulatory or signaling roles in cellular processes.
- Epigenetic factors: Molecules that influence gene expression and cellular phenotype without altering the DNA sequence, such as DNA methylation or histone modifications.
- Transposons: Mobile genetic elements capable of moving within a genome, potentially influencing gene expression and genomic rearrangements.
- Prions: Abnormal proteins that can induce the misfolding of normal proteins, leading to diseases known as prion diseases.
- Messenger RNA (mRNA): RNA molecules transcribed from DNA that carry the genetic information from the nucleus to the ribosomes for protein synthesis.
- Transfer RNA (tRNA): RNA molecules that carry specific amino acids to the ribosomes during protein synthesis.
- Ribosomal RNA (rRNA): RNA molecules that are structural components of ribosomes, the cellular machinery responsible for protein synthesis.
- Non-coding RNA (ncRNA): RNA molecules that do not code for proteins but have regulatory functions in gene expression and other cellular processes.
- Chromatin: The complex of DNA, histones, and other proteins that make up the chromosomes, regulating DNA accessibility and gene expression.
- Enhancers: DNA sequences that can enhance the transcription of genes by binding to specific transcription factors and promoting gene expression.
- Promoters: DNA sequences that initiate transcription by providing binding sites for RNA polymerase and other transcriptional machinery.
- Long non-coding RNAs (lncRNAs): RNA molecules longer than 200 nucleotides that do not code for proteins but participate in gene regulation and cellular processes.
- Telomeres: Repeated DNA sequences at the ends of chromosomes that protect them from degradation and maintain chromosomal stability.
- Telomerase: An enzyme that adds repetitive DNA sequences to telomeres, counteracting their shortening during DNA replication.
- Homologous recombination proteins: Proteins involved in the repair of DNA double-strand breaks using homologous DNA sequences as templates.
- DNA methyltransferases: Enzymes responsible for adding methyl groups to DNA, contributing to epigenetic modifications and gene regulation.
- Polycomb group proteins: Proteins involved in regulating gene expression by modifying chromatin structure and maintaining cellular identity.
- Cohesins: Protein complexes that help hold sister chromatids together during cell division and contribute to DNA repair and gene regulation.
- DNA helicases: Enzymes that unwind the DNA double helix during processes like DNA replication, repair, and transcription.
- DNA ligases: Enzymes that seal breaks in the DNA backbone during DNA replication, repair, and recombination.
- DNA repair proteins: Proteins involved in recognizing and repairing damaged DNA to maintain genome integrity.
- DNA methyl-binding proteins: Proteins that bind specifically to methylated DNA, playing a role in gene regulation and chromatin organization.
- RNA interference (RNAi) molecules: Small RNA molecules that can inhibit gene expression by targeting and degrading specific mRNA molecules.
- Non-coding circular RNAs (circRNAs): RNA molecules that form closed loops and are involved in gene regulation and cellular processes.
- DNA-binding proteins: Proteins that specifically bind to DNA sequences, regulating gene expression, replication, repair, and chromatin structure.
- Mitochondrial DNA (mtDNA): DNA found within mitochondria, responsible for encoding some of the genetic information necessary for mitochondrial function.
- RNA editing enzymes: Enzymes that catalyze changes in RNA sequences through post-transcriptional modifications, affecting RNA function.
- Telomere-binding proteins: Proteins that bind specifically to telomeric DNA, participating in telomere maintenance and genome stability.
- RNA helicases: Enzymes that unwind RNA molecules, contributing to RNA processing, translation, and other RNA-related processes.
- DNA topoisomerases: Enzymes that modify the topological structure of DNA by introducing or removing DNA supercoils.
- DNA methyl-demethylases: Enzymes that remove methyl groups from DNA, contributing to gene regulation and epigenetic modifications.
- Epigenetic modifiers (e.g., acetyltransferases, methyltransferases): Enzymes that add or remove chemical groups, such as acetyl or methyl groups, to DNA or histone proteins, influencing gene expression and chromatin structure.
- Non-histone chromosomal proteins: Proteins associated with chromosomes that are not histones, contributing to chromatin structure and gene regulation.
- Replication factors: Proteins involved in DNA replication, ensuring accurate and efficient duplication of the genome.
- Recombination enzymes: Enzymes involved in DNA recombination processes, such as homologous recombination or site-specific recombination.
- DNA proofreading and editing enzymes: Enzymes that ensure the accuracy of DNA replication by detecting and correcting errors in DNA sequences.
- Chromatin remodeling complexes: Protein complexes that use the energy of ATP hydrolysis to reposition or alter the structure of nucleosomes, regulating DNA accessibility and gene expression.
- RNA processing factors: Proteins involved in various steps of RNA processing, including splicing, capping, and polyadenylation.
- DNA damage sensors and repair regulators: Proteins that detect DNA damage and initiate signaling pathways to coordinate DNA repair processes.
- Transcriptional co-activators and co-repressors: Proteins that interact with transcription factors and the transcriptional machinery to enhance or suppress gene expression.
- DNA packaging proteins: Proteins that compact DNA into higher-order structures, facilitating its organization within the nucleus and regulating gene expression.
- Centromeric proteins: Proteins that play a role in the structure and function of centromeres, the regions of chromosomes essential for proper chromosome segregation during cell division.
- DNA replication origins and initiation factors: DNA sequences and proteins that facilitate the initiation of DNA replication at specific sites in the genome.
- DNA translocases: Enzymes that move along DNA strands, rearranging DNA structures, and facilitating processes such as DNA repair and recombination.
- Apoptosis-regulating proteins: Proteins involved in the regulation of programmed cell death (apoptosis), influencing cellular survival and development.
- DNA condensins: Protein complexes that aid in the condensation and compaction of DNA, particularly during mitosis and meiosis.
- Chromosomal segregation proteins: Proteins involved in the proper segregation of chromosomes during cell division, ensuring equal distribution of genetic material.
- DNA modification enzymes (e.g., DNA methyltransferases, demethylases): Enzymes that add or remove chemical modifications to DNA bases, affecting gene expression and epigenetic marks.
- DNA packaging proteins (e.g., H1 histones, protamines): Proteins that assist in further compacting DNA into higher-order structures, facilitating chromosome organization.
- DNA damage response proteins: Proteins involved in detecting DNA damage and activating signaling pathways that regulate DNA repair, cell cycle progression, and cell survival.
- DNA replication licensing factors: Proteins that control the initiation and licensing of DNA replication, ensuring that DNA is replicated only once per cell cycle.
- DNA-binding domains (e.g., zinc finger, helix-turn-helix): Structural motifs within proteins that enable specific binding to DNA sequences, regulating gene expression.
- RNA export and localization factors: Proteins involved in the transport of RNA molecules from the nucleus to the cytoplasm and their localization to specific cellular compartments.
- RNA stability and degradation factors: Proteins that regulate the stability and degradation of RNA molecules, influencing their abundance and half-life.
- RNA splicing and processing factors: Proteins involved in the splicing and processing of pre-mRNA molecules to generate mature, functional mRNA molecules.
- RNA-binding proteins: Proteins that bind to RNA molecules, influencing their localization, stability, translation, and other aspects of RNA metabolism.
- RNA chaperones: Proteins that assist in the proper folding and structural rearrangement of RNA molecules, ensuring their functional conformation.
- RNA granule components: Proteins that participate in the formation and regulation of RNA granules, which are dynamic structures involved in RNA localization, storage, and translation.
- DNA methylation readers and effectors: Proteins that specifically recognize and interact with methylated DNA, mediating downstream effects on gene expression and chromatin structure.
- RNA modifying enzymes (e.g., RNA methyltransferases): Enzymes that add chemical modifications, such as methyl groups, to RNA molecules, affecting their function and stability.
- DNA damage tolerance factors: Proteins involved in DNA damage tolerance mechanisms that allow DNA replication to continue despite lesions or roadblocks in the DNA template.
- RNA transport and localization factors: Proteins involved in the transport and localization of RNA molecules to specific subcellular compartments or sites of translation.
- RNA editing factors: Proteins that mediate RNA editing processes, modifying RNA sequences post-transcriptionally to alter their information content.
- DNA mismatch repair proteins: Proteins that recognize and repair mismatches or small insertion/deletion loops in DNA, ensuring replication fidelity and genome stability.
- RNA surveillance and quality control factors: Proteins that monitor and eliminate aberrant RNA molecules, ensuring the production of functional and high-quality RNA.
- DNA demethylases: Enzymes that activelyremove methyl groups from DNA, contributing to gene regulation and epigenetic modifications.
- RNA-binding proteins: Proteins that bind to RNA molecules, influencing their stability, localization, and function.
- DNA excision repair proteins: Proteins involved in excision repair mechanisms that remove damaged or incorrect nucleotides from DNA.
- Chromosomal remodeling factors: Proteins that modify the structure and organization of chromatin, influencing gene expression and DNA accessibility.
- DNA replication factors: Proteins necessary for DNA replication, ensuring accurate and efficient synthesis of the DNA molecule.
- DNA methylation modifiers: Enzymes or proteins that add or remove methyl groups from DNA, impacting gene expression and epigenetic modifications.
- Non-coding RNA-binding proteins: Proteins that specifically bind to non-coding RNA molecules, regulating their stability, localization, and function.
- Telomere-binding proteins: Proteins that bind to telomeres, protecting them from degradation, maintaining their structure, and regulating telomere length.
- DNA polymerases: Enzymes responsible for synthesizing DNA molecules during DNA replication and repair processes.
- Recombinases: Enzymes involved in DNA recombination processes, catalyzing the exchange of genetic material between DNA molecules.
- DNA-binding transcriptional regulators: Proteins that bind to specific DNA sequences and regulate gene expression by activating or repressing transcription.
- RNA-induced silencing complex (RISC): A multiprotein complex that includes small RNA molecules and targets specific mRNA molecules for degradation or translational repression.
- DNA methyltransferase inhibitors: Small molecules that inhibit the activity of DNA methyltransferases, influencing DNA methylation patterns and gene expression.
- Heterochromatin proteins: Proteins associated with heterochromatin, the condensed and transcriptionally inactive form of chromatin.
- Repair-associated factors: Proteins involved in DNA repair processes, including recognition, signaling, and execution of repair mechanisms.
- DNA replication licensing factors: Proteins that control the initiation of DNA replication and ensure that DNA is replicated only once per cell cycle.
- DNA damage response kinases: Enzymes that are activated in response to DNA damage and play a crucial role in coordinating DNA repair pathways and cell cycle checkpoints.
Note:
This comprehensive list that may encompasses a wide range of molecules involved in heredity, gene regulation, DNA repair, and various cellular processes. To aggregate/collect the data related with these areas and for the wellbeing of all of our loved one and all alives tha data allocated from various resources and to aggregate/collect and list the data we received help from AI applications and ChatGPT to collect and list this kind of various data.
Note that there are maybe hundreds or at least dozen hundreds more of these molecules. We’ll try to write, collect/aggregate and list other related data and list of molecules related with these.
Best wishes,
See also:
Some Related Articles:
https://fineupme.com/stem-cell-therapies-for-dogs/
https://fineupme.com/immortal-jellyfish-turritopsi-dohrnii/